Hierarchical Navigable Small Worlds (HNSW)
Hierarchical Navigable Small World (HNSW) is a graph-based algorithm that performs approximate nearest neighbor (ANN) searches in vector databases.
Read the entire series
- Introduction to Unstructured Data
- What is a Vector Database and how does it work: Implementation, Optimization & Scaling for Production Applications
- Understanding Vector Databases: Compare Vector Databases, Vector Search Libraries, and Vector Search Plugins
- Introduction to Milvus Vector Database
- Milvus Quickstart: Install Milvus Vector Database in 5 Minutes
- Introduction to Vector Similarity Search
- Everything You Need to Know about Vector Index Basics
- Scalar Quantization and Product Quantization
- Hierarchical Navigable Small Worlds (HNSW)
- Approximate Nearest Neighbors Oh Yeah (Annoy)
- Choosing the Right Vector Index for Your Project
- DiskANN and the Vamana Algorithm
- Safeguard Data Integrity: Backup and Recovery in Vector Databases
- Dense Vectors in AI: Maximizing Data Potential in Machine Learning
- Integrating Vector Databases with Cloud Computing: A Strategic Solution to Modern Data Challenges
- A Beginner's Guide to Implementing Vector Databases
- Maintaining Data Integrity in Vector Databases
- From Rows and Columns to Vectors: The Evolutionary Journey of Database Technologies
- Decoding Softmax Activation Function
- Harnessing Product Quantization for Memory Efficiency in Vector Databases
- How to Spot Search Performance Bottleneck in Vector Databases
- Ensuring High Availability of Vector Databases
- Mastering Locality Sensitive Hashing: A Comprehensive Tutorial and Use Cases
- Vector Library vs Vector Database: Which One is Right for You?
- Maximizing GPT 4.x's Potential Through Fine-Tuning Techniques
- Deploying Vector Databases in Multi-Cloud Environments
- An Introduction to Vector Embeddings: What They Are and How to Use Them
Latest Update: July 26
In the previous tutorial, we looked at scalar quantization and product quantization, two vector indexing strategies that reduce the overall size of the database without reducing the scope of our search. To better illustrate how scalar quantization and product quantization work, we also implemented our own versions in Python.
In this tutorial, we'll build on that knowledge by looking at today's most commonly used primary algorithm: Hierarchical Navigable Small Worlds (HNSW). Hierarchical Navigable Small World (HNSW) is a graph-based algorithm that performs approximate nearest neighbor searches (ANN) in vector databases. The HNSW algorithm performs ANN search very well in terms of speed and accuracy, making it an incredibly robust vector search algorithm. Unfortunately, despite being popular, understanding the HNSW algorithm can be tricky, but don't fret - in the next couple of sections, we'll break down HNSW into its steps, developing our own simple implementation along the way.
HNSW basics
Recall from a previous tutorial that there are four types of vector search indexes: hash-based, tree-based, cluster-based, and graph-based. HNSW fits firmly into the latter category: the graph index algorithm. It combines two core concepts—the probability skip list and Navigable Small World (NSW) graphs. Let's first dive into these two concepts individually before discussing the HNSW algorithm.
What are the probability skip lists?
First up: probability skip list. Recall the venerable linked list - a well-known data structure where each element maintains a pointer for the next element. Although linked lists work great for implementing LIFO and FIFO data structures such as stacks and queues, a major downside is their time complexity regarding random access: O(n). Skip lists aim to solve this problem by introducing additional layers, allowing for O(log n) random access time complexity by incurring extra memory (O(n log n) space complexity as opposed to O(n) for a normal linked list) and a bit of runtime overhead for inserts and deletes.
A probability skip list is a multi-level linked list where the upper levels maintain long connections. As we move down the layers, the connections become shorter and shorter, with the bottom layer being the "original" linked list containing all of the elements. The image below illustrates this probability skip list structure:
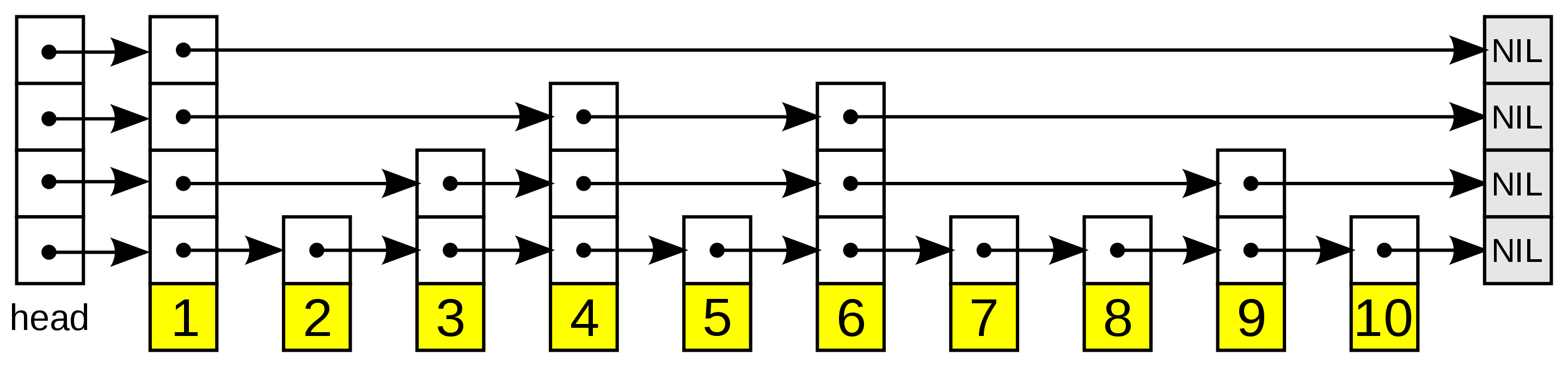 skip list structure
skip list structure
The skip list structure, illustrated. Higher layers have fewer elements.
We start at the highest layer to reach element i in a skip list. Once we find a node corresponding to an element in the list greater than i, we backtrack to the previous node and move to the layer below. This continues until we've found the element we're looking for. Note that skip lists only work for sorted lists, as we need a way to compare the magnitude of two objects directly.
Inserts work probabilistically. For any new element, we first need to figure out the layer with which the element appears first. The uppermost layer has the lowest probability, increasing probability as we move down in layers. The general rule is that any element in a layer will appear in layer above it with some pre-defined probability p. Therefore, if an element first appears in some layer l, it will also get added to layers l-1, l-2, and so on.
Note that, while it is possible to have a balanced skip list that performs no better than a standard linked list, the probability of this happening is incredibly low.
What is a Navigable Small World (NSW)?
Now that we've gotten skip lists out of the way, let's take some time to talk about Navigable Small Worlds (NSW). NSW is a graph-based algorithm that finds approximate nearest neighbors in a dataset. The general idea here is first to imagine many nodes in a network. Each node will have short-, medium-, and long-range connections to other nodes. When performing a vector search, we'll begin at some pre-defined entry point. From there, we'll evaluate connections to other nodes and jump to the one closest to the one we hope to find. This process repeats until we've found our nearest neighbor.
This type of search is called greedy search. This algorithm works for small NSWs in the hundreds or thousands of nodes, but it tends to break down for much larger NSWs. We can fix this by increasing the average number of short-, medium-, and long-range connections for each node, but this increases the overall complexity of the network and results in longer search times. In the absolute "worst" case, where each node is connected to every other node in our dataset, NSW is no better than naïve (linear) search.
NSWs are cool, but how does this relate to vector search? The idea here is to imagine all vectors in our dataset as points in an NSW, with long-range connections defined by vectors dissimilar from one another and the opposite for short-range connections. Recall that vector similarity scores are measured with a similarity metric - typically L2 distance (also called Euclidean distance) or inner product (IP) for floating point vectors and Jaccard or Hamming distance for binary vectors.
By constructing a navigable small world graph (NSW graph) with dataset vectors as vertices, we can effectively perform nearest neighbor search by greedily traversing the NSW towards vertices closer and closer to our query vector.
What is HNSW (Hierarchical Navigable Small Worlds)?
When it comes to vector search or vector similarity search, we often have dataset sizes of hundreds of millions or even billions of vectors. Plain NSWs are less effective at this scale, so we'll need a better graph structure.
 better_graph.png
better_graph.png
Enter HNSW.
HNSW extends NSW by borrowing from the concept of skip lists. Like the skip list, HNSW maintains multiple layers (hence the term Hierarchical Navigable Small World), only of NSWs instead of linked lists. The uppermost layer of an HNSW graph has few nodes and the longest links, while the bottommost layer has all nodes and the shortest links. During the search process, we enter a pre-defined point in the uppermost layer and greedily route ourselves toward the nearest neighbor to our query vector. Once we reach the nearest node, we repeat this process to the second layer of the HNSW graph. This continues until we've reached our nearest neighbor.
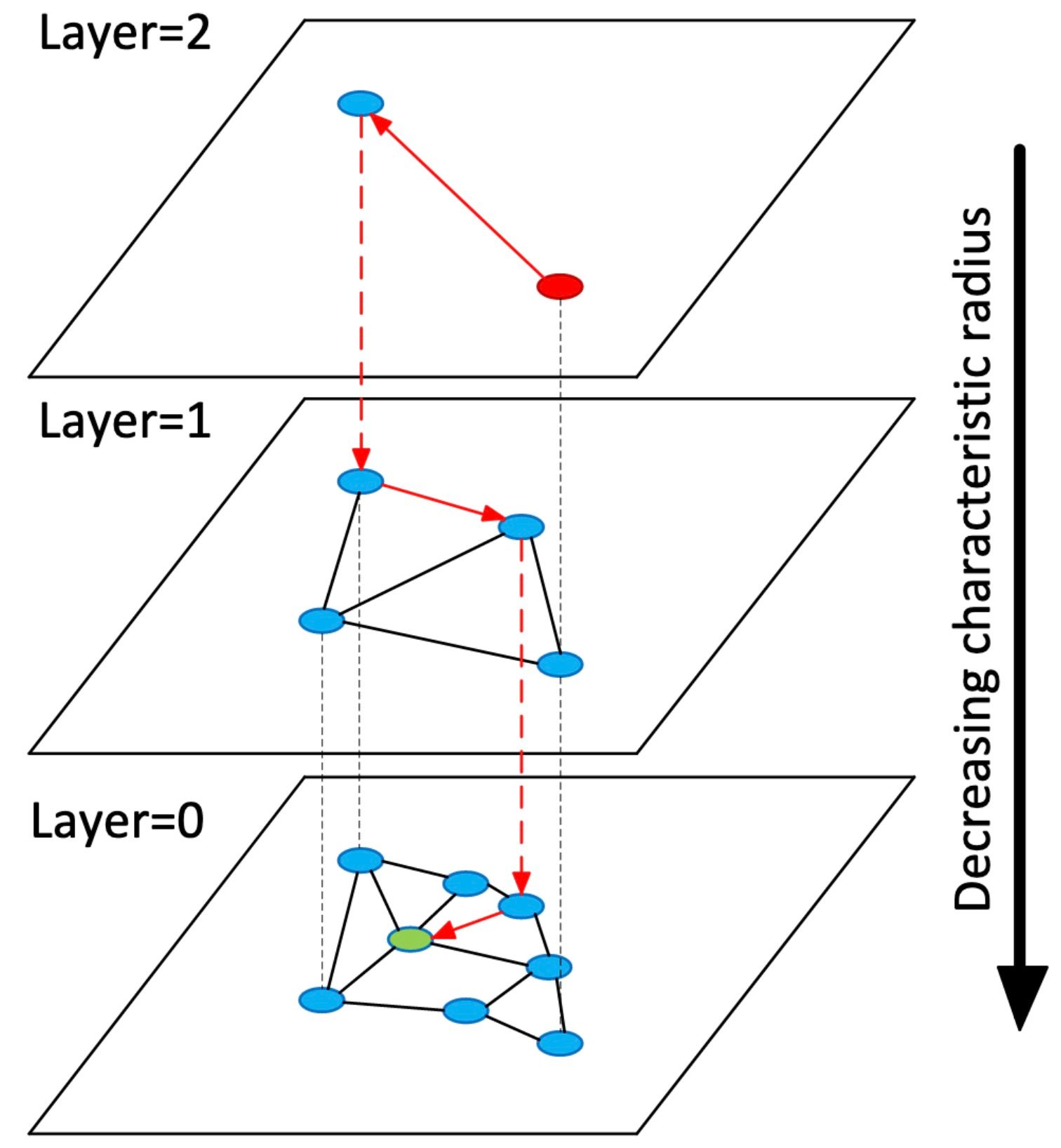 hnsw_visualized_hnsw graph.jpg
hnsw_visualized_hnsw graph.jpg
A diagram from the HNSW paper which visualizes the layered graph concept. From https://arxiv.org/abs/1603.09320.
Inserts work similarly to the skip list. For some vector v, We first traverse the first layer of the graph, finding its nearest neighbor before moving to the layer below it. Then, we traverse the graph again to find its nearest neighbor in the second layer. This process until we've reached the nearest neighbor in the bottommost graph.
We must determine which links (connections between vertices) to create from here. Again, we have a pre-defined parameter M which determines the maximum number of bidirectional links we can add. These links are usually set as the nearest neighbors to v, but other heuristics can also be used. The same process repeats for the upper layers, assuming the vector appears.
As with the skip list, the query vector will appear in upper layers with exponentially decreasing probability. Specifically, the HNSW paper uses the equation floor(-ln(rand(0, 1))), where rand(0, 1) is a random number sampled from a uniform distribution between (0, 1]. Note how this does not constrain the minimum distance between any two vertices/vectors in a particular layer - we may end up with a poorly constructed graph. Still, the probability of this happening is incredibly low, especially as we scale up the number of vectors in the HNSW index.
How to Implement HNSW
HNSW is not trivial, so we'll implement only a basic version here. As usual, let's start with creating a dataset of (128 dimensional) vectors:
>>> import numpy as np
>>> dataset = np.random.normal(size=(1000, 128))
The first step is to build the HNSW index. We'll need to add each vector to our dataset to do so. So let's first create a data structure to hold our index. In this basic example, we'll use a list of lists to represent the index, with the inner lists corresponding to each layer/graph:
>>> L = 5 # 5-layer HNSW
>>> index = [[] for _ in range(L)]
Every element in each graph is a 3-tuple containing the vector, a list of indexes the vector links to within the graph, and the index for the corresponding node in the layer below it. For the bottommost layer, the third element of the 3-tuple will be set to None.
Since every insert first requires a search for the nearest neighbor in graph, let's implement that first. We can traverse any of the subgraphs in the index as so:
def _search_layer(graph, entry, query, ef=1):
best = (np.linalg.norm(graph[entry][0] - query), entry)
nns = [best]
visit = set(best) # set of visited nodes
candid = [best] # candidate nodes to insert into nearest neighbors
heapify(candid)
# find top-k nearest neighbors
while candid:
cv = heappop(candid)
if nns[-1][0] < cv[0]:
break
# loop through all nearest neighbors to the candidate vector
for e in graph[cv[1]][1]:
d = np.linalg.norm(graph[e][0] - query)
if (d, e) not in visit:
visit.add((d, e))
# push only "better" vectors into candidate heap
if d < nns[-1][0] or len(nns) < ef:
heappush(candid, (d, e))
insort(nns, (d, e))
if len(nns) > ef:
nns.pop()
return nns
This code snippet is more involved, but it's much easier to understand with some explanation. Here, we use a heap to implement a priority queue, which we use to order nearest neighbor vectors in the graph. Like all of the previous examples, I'm using L2 distance (Euclidean Distance) here, but this code can also be extended to other distance metrics. We first populate the heap with the entry point.
Here, all we're doing is implementing greedy search. At every iteration, we aim to update two variables: nns, our output list of nearest neighbors, and candid, a heap of candidate points. So, first, we evaluate all nearest neighbors to the "best" vector in candid, adding better (better means closer to the query vector) vectors to the output list of nearest neighbors and the heap of candidate points for evaluation on the next iteration. This repeats until one of two stopping conditions is reached: we either run out of candidate points to evaluate, or we've determined that we can no longer do any better than we already have.
With top-k graph search out of the way, we can now implement the top-level search function for searching the entire HNSW index:
def search(index, query, ef=1):
# if the index is empty, return an empty list
if not index[0]:
return []
best_v = 0 # set the initial best vertex to the entry point
for graph in index:
best_d, best_v = _search_layer(graph, best_v, query, ef=1)[0]
if graph[best_v][2]:
best_v = graph[best_v][2]
else:
return _search_layer(graph, best_v, query, ef=ef)
We first start at the entry point (zeroth element in the uppermost graph), and search for the nearest neighbor in each index layer until we reach the bottommost layer. Recall that the final element of the 3-tuple will resolve to None if we are at the bottommost layer - this is what the final if statement is for. Once we reach the lowermost layer, we search the graph using best_v as the entry point.
Let's go back go the HNSW insert. We'll first need to figure out which layer to insert our new vector into. This is fairly straightforward:
def _get_insert_layer(L, mL):
# ml is a multiplicative factor used to normalized the distribution
l = -int(np.log(np.random.random()) * mL)
return min(l, L)
With everything in place, we can now implement the insertion function.
def insert(self, vec, efc=10):
# if the index is empty, insert the vector into all layers and return
if not index[0]:
i = None
for graph in index[::-1]:
graph.append((vec, [], i))
i = 0
return
l = _get_insert_layer(1/np.log(L))
start_v = 0
for n, graph in enumerate(index):
# perform insertion for layers [l, L) only
if n < l:
_, start_v = _search_layer(graph, start_v, vec, ef=1)[0]
else:
node = (vec, [], len(_index[n+1]) if n < L-1 else None)
nns = _search_layer(graph, start_v, vec, ef=efc)
for nn in nns:
node[1].append(nn[1]) # outbound connections to NNs
graph[nn[1]][1].append(len(graph)) # inbound connections to node
graph.append(node)
# set the starting vertex to the nearest neighbor in the next layer
start_v = graph[start_v][2]
If the index is empty, we'll insert vec into all layers and return immediately. This serves to initialize the index and allow for successful insertions later. If the index has already been populated, we begin insertion by first computing the insertion layer via the get_insert_layer function we implemented in the previous step. From there, we find the nearest neighbor to the vector in the uppermost graph. This process continues for the layers below it until we reach layer l, the insertion layer.
For layer l and all those below it, we first find the nearest neighbors to vec up to a pre-determined number ef. We then create connections from the node to its nearest neighbors and vice versa. Note that a proper implementation should also have a pruning technique to prevent early vectors from being connected to too many others - I'll leave that as an exercise for the reader :sunny:.
We now have both search (query) and insert functionality complete. Let's combine everything in a class:
from bisect import insort
from heapq import heapify, heappop, heappush
import numpy as np
from ._base import _BaseIndex
class HNSW(_BaseIndex):
def __init__(self, L=5, mL=0.62, efc=10):
self._L = L
self._mL = mL
self._efc = efc
self._index = [[] for _ in range(L)]
@staticmethod
def _search_layer(graph, entry, query, ef=1):
best = (np.linalg.norm(graph[entry][0] - query), entry)
nns = [best]
visit = set(best) # set of visited nodes
candid = [best] # candidate nodes to insert into nearest neighbors
heapify(candid)
# find top-k nearest neighbors
while candid:
cv = heappop(candid)
if nns[-1][0] > cv[0]:
break
# loop through all nearest neighbors to the candidate vector
for e in graph[cv[1]][1]:
d = np.linalg.norm(graph[e][0] - query)
if (d, e) not in visit:
visit.add((d, e))
# push only "better" vectors into candidate heap
if d < nns[-1][0] or len(nns) < ef:
heappush(candid, (d, e))
insort(nns, (d, e))
if len(nns) > ef:
nns.pop()
return nns
def create(self, dataset):
for v in dataset:
self.insert(v)
def search(self, query, ef=1):
# if the index is empty, return an empty list
if not self._index[0]:
return []
best_v = 0 # set the initial best vertex to the entry point
for graph in self._index:
best_d, best_v = HNSW._search_layer(graph, best_v, query, ef=1)[0]
if graph[best_v][2]:
best_v = graph[best_v][2]
else:
return HNSW._search_layer(graph, best_v, query, ef=ef)
def _get_insert_layer(self):
# ml is a multiplicative factor used to normalize the distribution
l = -int(np.log(np.random.random()) * self._mL)
return min(l, self._L-1)
def insert(self, vec, efc=10):
# if the index is empty, insert the vector into all layers and return
if not self._index[0]:
i = None
for graph in self._index[::-1]:
graph.append((vec, [], i))
i = 0
return
l = self._get_insert_layer()
start_v = 0
for n, graph in enumerate(self._index):
# perform insertion for layers [l, L) only
if n < l:
_, start_v = self._search_layer(graph, start_v, vec, ef=1)[0]
else:
node = (vec, [], len(self._index[n+1]) if n < self._L-1 else None)
nns = self._search_layer(graph, start_v, vec, ef=efc)
for nn in nns:
node[1].append(nn[1]) # outbound connections to NNs
graph[nn[1]][1].append(len(graph)) # inbound connections to node
graph.append(node)
# set the starting vertex to the nearest neighbor in the next layer
start_v = graph[start_v][2]
Boom, done!
A Summary of HNSW
In this tutorial, we explored Hierarchical Navigable Small Worlds (HNSW), a powerful graph-based vector similarity search strategy that involves multiple layers of connected graphs. This HNSW algorithm is often the go-to choice for Milvus users looking for maximum query performance.
In the next tutorial, we'll continue our deep dive into indexing strategies with Approximate Nearest Neighbor Oh Yeah (ANNOY) - a tree-based indexing algorithm that I particularly enjoy because of its playful name.
Take another look at the Vector Database 101 courses
- Introduction to Unstructured Data
- What is a Vector Database?
- Comparing Vector Databases, Vector Search Libraries, and Vector Search Plugins
- Introduction to Milvus
- Milvus Quickstart
- Introduction to Vector Similarity Search
- Vector Index Basics and the Inverted File Index
- Scalar Quantization and Product Quantization
- Hierarchical Navigable Small Worlds (HNSW)
- Approximate Nearest Neighbors Oh Yeah (ANNOY)
- Choosing the Right Vector Index for Your Project
- DiskANN and the Vamana Algorithm
Webinars & Events You May Be Interested in
- Zilliz Webinar | Vector Database 101: A Crash Course
- Zilliz Webinar | Fireside chat with HNSW author Yury Malkov
- Zilliz Webinar | Vector Search Best Practices
- Zilliz Webinar | Tutorial: Working with LLMs at Scale
- HNSW basics
- What is HNSW (Hierarchical Navigable Small Worlds)?
- How to Implement HNSW
- A Summary of HNSW
- Take another look at the Vector Database 101 courses
- Webinars & Events You May Be Interested in
Content
Start Free, Scale Easily
Try the fully-managed vector database built for your GenAI applications.
Try Zilliz Cloud for FreeKeep Reading

Introduction to Unstructured Data
Buckle up for the first tutorial in our Vector Database 101 series and untangle the intricacy around Milvus with us every week.

Decoding Softmax Activation Function
This article will discuss the Softmax Activation Function, its applications, challenges, and tips for better performance.

Deploying Vector Databases in Multi-Cloud Environments
Multi-cloud deployment has become increasingly popular for services looking for as much uptime as possible, with organizations leveraging multiple cloud providers to optimize performance, reliability, and cost-efficiency.
